Abstract
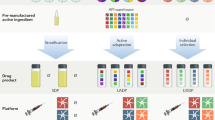
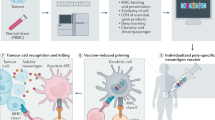
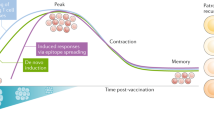
The study of tumour-specific antigens (TSAs) as targets for antitumour therapies has accelerated within the past decade. The most commonly studied class of TSAs are those derived from non-synonymous single-nucleotide variants (SNVs), or SNV neoantigens. However, to increase the repertoire of available therapeutic TSA targets, ‘alternative TSAs’, defined here as high-specificity tumour antigens arising from non-SNV genomic sources, have recently been evaluated. Among these alternative TSAs are antigens derived from mutational frameshifts, splice variants, gene fusions, endogenous retroelements and other processes. Unlike the patient-specific nature of SNV neoantigens, some alternative TSAs may have the advantage of being widely shared by multiple tumours, allowing for universal, off-the-shelf therapies. In this Opinion article, we will outline the biology, available computational tools, preclinical and/or clinical studies and relevant cancers for each alternative TSA class, as well as discuss both current challenges preventing the therapeutic application of alternative TSAs and potential solutions to aid in their clinical translation.
Similar content being viewed by others
Introduction
The role of tumour-specific antigens (TSAs) as targets of anticancer immunity was first recognized in the last century, with studies of TSA-based vaccines becoming more prevalent in the past decade1,2,3 (Box 1). Neoantigens are defined here as a subset of TSAs generated by non-synonymous mutations and other genetic variations specific to the genome of a tumour, presented by major histocompatibility complex (MHC) molecules, and recognized by endogenous T cells. The most commonly studied class of neoantigens are those derived from single-nucleotide variants (SNVs), which cause non-synonymous changes in a protein that subsequently may trigger antigen-specific T cell responses against the tumour. These conventional neoantigens have the distinct advantage over other classes of tumour antigens (for example, tumour-associated antigens and cancer–testis antigens) of having no expression in normal tissues4. As a result, T cells with specificity for these neoantigens can escape negative selection in the thymus, leading to the generation of a TSA-specific T cell repertoire5.
Despite the advantages of SNV neoantigens, their applicability as vaccine targets may be limited to cancers with highly immunogenic neoantigens, likely a subset of the total neoantigen load for any given tumour. Metastatic melanoma (which contains the highest SNV burden of any cancer6) has been the primary focus of initial neoantigen clinical studies2,3, and in this tumour type, as in lung cancer, tumour mutational burden (which estimates neoantigen load) has been associated with the response to immune checkpoint inhibition7. One hypothesis for this association is the increased likelihood in these tumour types of neoantigen generation and T cells bearing neoantigen-specific T cell receptors (TCRs). However, the number of neoantigens required for driving a clinical response is unknown, and it has been shown that tumours with a low mutational burden can have neoantigen-specific T cell populations boosted by therapeutic personalized neoantigen vaccines8,9.
Many investigators, including our group, have begun to evaluate alternative TSAs — defined as high-specificity tumour antigens arising from non-SNV-containing genomic sources. Unlike SNV neoantigens, alternative TSAs are not necessarily restricted to protein-coding exons, allowing for a greater repertoire of available targets. Predicted tumour antigen burden has demonstrated that the expression of various classes of TSAs is not always correlated, with some SNV-low cancers containing high alternative-TSA expression. This is exemplified by clear-cell renal-cell carcinoma (ccRCC), an immune checkpoint inhibitor sensitive cancer that has a low predicted SNV burden but high expression of predicted frameshift neoantigens10 and tumour-specific endogenous retroviral antigens11. Thus, studying these alternative TSAs may broaden the scope and increase the number of targets available to test in therapeutic vaccines and/or cellular therapies. Additionally, leukaemia and sarcoma (which are among the cancers with the lowest predicted SNV burden12) express gene fusions13,14 and splice variant transcripts15,16 shared across multiple tumours, potentially allowing for universal off-the-shelf therapies.
In this Opinion article, we will characterize several major classes of alternative TSAs, including those generated from mutational frameshifts, splice variants, gene fusions, endogenous retroelements and other classes, such as human leukocyte antigen (HLA)-somatic mutation-derived antigens and post-translational TSAs (Fig. 1; Table 1). One class of TSA not covered here is the viral-derived cancer antigens (for example, human papillomavirus (HPV) and Epstein–Barr virus (EBV)), which have been previously reviewed17,18,19,20,21. We will begin by providing a brief overview of TSA computational prediction and then discuss the biology, available computational tools, preclinical and/or clinical studies, and relevant cancers for each alternative TSA class. Finally, we will discuss the current challenges impeding therapeutic application of alternative TSAs and solutions to aid their clinical translation. In addition to a review of the literature, recent studies (including several from our group) have provided estimates of the antigenic burden of each TSA class among The Cancer Genome Atlas (TCGA) pan-cancer data (including selected tumour-specific viral antigens), which we have compiled here as a resource (Fig. 2 and Supplementary Fig. 1; viral antigens including those derived from EBV, herpes simplex virus and cytomegalovirus are included for comparison).
Mutations and other tumour-specific nucleotide sequences (shown in red) can be observed at the genomic DNA level, where they undergo transcription (1) and splicing to form mRNA (2). Alternative splicing can occur at this step, to form splice variant mRNA. Next, translation occurs on variant mRNA, resulting in the production of variant proteins (3). Post-transcriptional frameshifts (for example, ribosomal slippage, among other mechanisms) can occur at this step, resulting in frameshifted protein variants. These proteins can then undergo proteasomal degradation (4) and transport to the endoplasmic reticulum (ER), to subsequently be loaded on major histocompatibility complexes (MHCs) (5). Other forms of post-translational frameshift can occur during these steps (for example, protein splicing). Finally, peptides containing variant sequences can be presented at the cell surface in the context of MHC, resulting in T cell targetable tumour-specific antigens (6). Chr, chromosome; ERV, endogenous retrovirus; INDEL, insertion or deletion; SNV, single-nucleotide variant.
Plots represent the number of unique identified epitopes by The Cancer Genome Atlas (TCGA) cancer type. Insertion or deletion (INDEL) neoantigen counts have demonstrated significant correlation with single-nucleotide variant (SNV) neoantigens among all cancer types (coefficient: 0.81, P < 0.0001). Notable outliers in this correlation are kidney renal clear-cell carcinoma (KIRC; commonly known as clear-cell renal-cell carcinoma (ccRCC)) and kidney renal papillary cell carcinoma (KIRP; commonly known as papillary RCC), where the INDEL-to-SNV ratio is significantly higher than in other cancer types (ccRCC, 0.85, and papillary RCC, 0.90; all others, 0.43 – 0.72). Analysis of splice variant antigens has demonstrated a burden similar to that of INDEL neoantigens, with significant correlations with both INDEL and SNV neoantigen burden. A notable outlier is thyroid cancer (thyroid carcinoma (THCA)), where the average number of splice variant antigens per sample is higher than that of SNV neoantigens. The mean burden of fusion-derived neoantigens is highest in sarcomas (for example, sarcoma (SARC), 1.1; uterine carcinosarcoma (UCS), 0.78), with the carcinoma fusion burden being highest in breast (breast invasive carcinoma (BRCA), 0.70) and prostate (prostate adenocarcinoma (PRAD), 0.58) cancer. Testicular cancer (testicular germ cell tumour (TGCT)) has a substantially greater burden of human endogenous retrovirus (hERV)-derived tumour-specific antigens (TSAs) than any other TCGA cancer type. SNV and INDEL epitopes are derived from Thorsson et al.12. Fusion epitopes are derived from Gao et al.199. Splice variant epitopes are derived from Jayasinghe et al.52. The viral epitopes are derived from Selitsky et al.200; the hERV epitopes are derived from differentially expressed hERVs (>10-fold tumour-versus-mean normal expression by DESeq2) in Smith et al.11; all TSA classes represent the average numbers of predicted class I human leukocyte antigen (HLA) binders (8–11 mers, <500 nM) predicted from NetMHCPan. Stomach adenocarcinoma (STAD) INDEL and SNV calls were absent from Thorsson et al.12, and oesophageal carcinoma, acute myeloid leukaemia and ovarian serous cystadenocarcinoma were not included in all original reports and so are absent from the figure. The data shown represent reanalysis of the above reports, with modification of the data to derive values comparable across TSA groups. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; CESC, cervical and endocervical cancers; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, lymphoid neoplasm diffuse large B cell lymphoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; LGG, brain lower-grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; PAAD, pancreatic adenocarcinoma; PCPG, phaeochromocytoma and paraganglioma; READ, rectum adenocarcinoma; SKCM, skin cutaneous melanoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UVM, uveal melanoma. A version of these data with individual numbers of unique TSAs by cancer type is available online (Supplementary Fig. 1).
Computational prediction of TSAs
Recent advancements in DNA and RNA sequencing (RNA-seq) have enabled the development of genomic and computational methods of TSA prediction (Table 2). Methods for generating TSA immunotherapies generally rely on a conserved set of steps: variant calling, HLA typing, peptide enumeration, HLA binding prediction and therapy generation (Fig. 3). Variant calling is the identification of genomic regions with tumour specificity. In the case of SNVs, insertion or deletion (INDEL) mutations and gene fusions, variants are derived from mutations within the tumour exome that are not expressed by germline DNA. In contrast, endogenous retroelement-derived antigens are identified from RNA expression data, selecting for elements with higher expression in the tumour than in matched normal tissues. Splice variant antigens can be identified through a variety of techniques, discussed in-depth later. Subsequently, tumour HLA typing is derived using an HLA caller (for example, POLYSOLVER22, OptiType23, PHLAT24, HLAScan25 or HLAProfiler26), which relies on DNA and/or RNA-seq data, depending upon the platform. Peptide enumeration is then performed, whereby variant genomic regions are translated into peptide sequences, with removal of translation-incompatible sequences such as nonsense mutations. Following this, HLA binding prediction is performed using prediction software (for example, NetMHCpan27), with higher-affinity peptides being characterized by either ranked percentiles or KD values of ≤500 nM (the commonly accepted binding affinity cutoff in the field)3,27,28. The majority of MHC binding affinity prediction tools rely on machine-learning algorithms (including artificial neural networks) trained on validated epitope reference databases, where peptide binding to MHC molecules has been measured using biochemical assays29,30. Finally, the predicted TSAs are used to generate a therapeutic product, either as a vaccine (that is, a DNA or RNA, peptide or dendritic cell vaccine) or a cellular therapy product (that is, adoptive T cell therapy). Below, we will discuss the biology of each alternative TSA class, with detailed descriptions of the available computational prediction tools.
a | The identification of tumour-specific antigens begins with variant calling. This can be done through the comparison of tumour versus normal-tissue DNA sequences (single-nucleotide variants (SNVs) and insertions or deletions (INDELs)) or RNA sequences (splice variants, fusions, viral sequences and retroelements) to look for tumour-specific variants in the exome or tumour-specific transcripts in the transcriptome, respectively. b | Tumour human leukocyte antigen (HLA) typing is performed to enable downstream major histocompatibility complex (MHC) binding prediction. c | Peptide enumeration occurs through the translation of variant nucleotide sequences into their respective amino acid sequences, filtering for translation-incompatible sequences, such as those containing intervening stop codons or those with low evidence of RNA expression. These polypeptides are then used to derive 8–11 mer sequences (for MHC class I epitopes) or 15 mer sequences (MHC class II epitopes). d | This then enables downstream MHC or HLA binding prediction of each sequence. Binders are typically defined in the literature as those with a predicted binding affinity (KD) of ≤500 nM or are selected from those with the highest ranked percentile for predicted binding affinity. Other filtering criteria may be performed after this step, such as immunogenicity prediction or filtering away sequences with high homology to self-antigens. e | Finally, therapies are generated using predicted tumour-specific antigens. These can be DNA, RNA or peptide vaccines or cellular therapies such as adoptive T cell therapy.
Mutational frameshift neoantigens
Biology of INDEL mutations
INDEL mutations are derived from the insertion of base pairs into or deletion from the genome, which has the capacity to generate non-synonymous novel open reading frames, known as mutational frameshifts. INDEL-derived neoantigens have been hypothesized (but not yet proven) to generate more robust immune responses than SNV-derived neoantigens, as their sequences are completely unique from germline sequences downstream of the INDEL10,31. Epitopes generated from these mutations could induce a T cell response similar to SNV neoantigens, due to decreased potential for negative selection in the thymus against the INDEL neoantigen-specific T cell.
Cancer types that are particularly relevant for targeting INDEL neoantigens include microsatellite instability-high (MSI-H) tumours, as well as all renal cell carcinomas (RCCs). Early studies examining the role of INDEL mutations for antitumour immunity were mainly pursued in colon cancer, where MSI caused by hereditary diseases (for example, Lynch syndrome) and in sporadic tumours (MSI-H in 15%) is common32,33. MSI-H tumours are also observed in other non-hereditary cancers, including gastric, endometrial and pancreatic cancers34. MSI-H cancers are characterized by impaired DNA mismatch repair pathways and are associated with significantly greater INDEL burden than non-MSI-H tumours31,35. The association between INDEL burden and the presence of tumour-infiltrating T cells has been well described in the literature, providing early support for the hypothesis that MSI-H tumours would be susceptible to immunotherapies31,36,37,38,39. Concurrent with these findings, immune checkpoint inhibitors have demonstrated clinical activity for patients with MSI-H tumours, independent of the tissue of origin40. As a result, MSI-H tumours are the only non-tissue-restricted class of tumours with US Food and Drug Administration (FDA) approval for immune checkpoint inhibitor therapy41. In MSI-H tumours, the burden of both SNV and INDEL neoantigens is high, making both neoantigen classes potentially useful for targeted therapy10.
In contrast, RCC contains relatively few SNVs, despite having immune infiltrates and a high clinical response rate to immune checkpoint inhibitor therapy42. A potential explanation for this was explored recently by Turajlic et al.10, whereby examining the pan-cancer INDEL profile in The Cancer Genome Atlas (TCGA) dataset revealed that all RCC subtypes (clear-cell RCC (ccRCC), renal papillary cell carcinoma and chromophobe RCC) have the highest proportion and number of INDEL mutations of any cancer types. The presence of INDELs was also associated with immune features (for example, T cell activation and immune checkpoint inhibitor response) in three individual cohorts of patients with melanoma. While the number of predicted INDELs across the pan-cancer cohort was orders of magnitude lower than the SNV mutations, they were estimated to produce approximately 3 to 9 times more predicted neoantigens per mutation than SNVs10.
Tools for predicting INDEL-derived neoantigens
Currently, we are aware of at least six tools in peer-reviewed publications with the capacity to predict INDEL-derived neoantigens — pVACseq43, Neopepsee44, MuPeXI45, Epidisco46, Antigen.garnish47, and TSNAD48 (not included here are the custom neoantigen prediction pipelines being used in translational and clinical studies, which may contain proprietary methods for antigen prediction along with integration of a publicly available variant caller (for example, Indelocator and Strelka49) and peptide–MHC binding prediction methods). Among these tools, Neopepsee is unique in its integration of machine-learning algorithms to predict immunogenicity — as well as peptide–MHC binding — a feature not easily validated biologically in human neoantigen studies before the induction of therapy.
Translation of INDEL-derived antigens into the clinic
A rare example of a publicly shared neoantigen has been observed in a common frameshift mutation in the gene transforming growth factor-β receptor type 2 (TGFBR2), frequently found in Lynch syndrome and 15% of sporadic gastric and colon cancers with MSI39. Three independent studies published in 2001 demonstrated HLA-specific epitopes generated from mutated TGFβR2 capable of generating antigen-specific T cells, one associated with MHC class I–CD8+ T cell responses31,39 and one with MHC class II–CD4+ T cell responses50. A more recent study from Inderberg et al.51 isolated cytotoxic T lymphocytes (CTLs) from a patient with colon cancer who had shown greater than 10-year survival after vaccination with a TGFβR2 frameshift mutation-derived peptide, and used these CTLs to generate a TCR-paired α- and β-chain clone, which was subsequently transfected into both CD4+ and CD8+ T cells. The transfected T cells demonstrated evidence of efficacy against colon cancer cell lines containing the TGFβR2 mutation both in vitro (cytotoxicity and cytokine release) and in vivo (an immunodeficient xenograft mouse model).
A recent publication from Ott et al.3 studied the use of personalized neoantigen vaccines in the treatment of metastatic melanoma, with prioritization of INDEL neoantigens in their prediction pipeline. Four unique INDEL mutations across six tumours were predicted, with T cell cultures generated that were specific to two of those INDEL neoantigens (one CD4+ T cell epitope and one CD8+ T cell epitope), which in turn demonstrated detectable interferon-γ (IFNγ) secretion in response to their respective epitopes. This was compared with only 3–5 of 28 predicted SNV neoantigens, which exhibited IFNγ responses of a similar concentration. Although INDEL neoantigen cross-reactivity with the respective reference wild-type epitope was not measured (presumably as it was expected there would be no cross-reactivity), over half of the SNV-specific T cells demonstrated cross-reactivity with their wild-type epitope at escalating concentrations. Due to the small patient cohort and follow-up so far only at 20–32 months, the clinical benefit of INDEL neoantigens cannot yet be determined from this study.
Splice variant antigens
Splice variant antigen frequency in cancer
Splice variant antigens are post-transcriptionally derived TSAs arising from alternative splicing events, including those from mRNA splice junction mutations52,53,54,55,56,57, intron retention58,59,60,61,62,63 or dysregulation of the spliceosome machinery in the tumour cell15,64,65. Other types of post-transcriptionally derived TSAs include alternative ribosomal products (for example, ribosomal frameshifting66,67, non-canonical initiation68,69,70,71, termination codon read-through69, reverse-stand transcription72 and doublet decoding73) and post-translational splicing74,75,76 — these two mechanisms are difficult to apply in anticancer therapies, given the lack of tools for predicting such products.
The study of splice variant proteins has historically focused on haematological malignancies, with splice variant protein expression being understudied in solid tumours. As such, putative splice variant antigens derived from these proteins have received less attention in solid tumours, with expression only recently validated77. In haematological cancers in which SNV burden is relatively low6, splice variant antigens could broaden the number of available TSA targets for therapeutic application. Splice variant proteins can arise through cis-acting mutations that disrupt or create splice site motifs or through trans-acting alterations in slicing factors that have historically been identified in haematological malignancies77,78. The role of spliceosome machinery in the generation of splice variants in haematological malignancies is a current area of investigation. Mutations in spliceosome proteins (for example, splicing factor 3b subunit 1 (SF3B1), serine- and arginine-rich splicing factor 2 (SRSF2), U2 small nuclear RNA auxiliary factor 1 (U2AF1) and U2AF2) are common in myelodysplastic syndrome, acute myeloid leukaemia (AML), chronic myelomonocytic leukaemia (CMML), and chronic lymphocytic leukaemia (CLL)79,80,81,82,83. Sharing of these spliceosome protein mutations across haematological cancer types has led to the hypothesis that spliceosome dysregulation may cause the expression of splice variant mRNAs, which are not detectable in normal tissues, leading to the translation of TSAs84,85,86. Beyond haematological malignancies, recent reanalysis of the TCGA pan-cancer dataset demonstrated a strong association between somatic mutations in components of the spliceosome machinery and the expression of splice variant products77, providing evidence for the relevance of splice variant antigens in solid tumours.
Tools for predicting splicing events and splice variant antigens
Several types of splice variant callers have been described in the literature. Two of these tools, Spliceman87 and MutPred Splice88, predict the capacity of exonic variants surrounding an annotated splice junction to interfere with normal splicing. Other tools provide de novo identification of alternative splicing events, including JuncBase89, SpliceGrapher90, rMATS91, SplAdder92 and ASGAL93. Many of these tools (for example, SpliceGrapher, SplAdder and ASGAL) predict alternative splicing events through the generation of splicing graphs. This splicing graph is generated through comparisons of spliced alignments of RNA-seq reads against a genome reference, which consists of vertices (nodes) that represent predicted splicing sites for a given gene as well as edges that represent exons and introns between splicing sites. In addition to these splice variant callers, at least one peer-reviewed tool, Epidisco46 (the computational pipeline for the multi-institutional PGV-001 personalized vaccine trial94), has been described with the capacity to predict for splice variant antigens.
Jayasinghe et al.52 reported MiSplice, which integrates DNA-seq and RNA-seq data in order to discover mutation-induced splice sites, which they applied to the TCGA pan-cancer dataset. Splice variant mutations contained 2–2.5x more predicted TSA candidates than did SNVs, with some tumorigenesis-related genes containing ≥40 unique predicted TSAs. Furthermore, predicted splice variant antigen burden was correlated with programmed cell death 1 ligand 1 (PDL1) expression, suggesting that PDL1 blockade therapy may be efficacious in tumours with a high frequency of splice variant antigens. Additionally, Kahles et al.77 reported a comprehensive analysis of splice variants in the TCGA pan-cancer dataset and then used mass spectrometry to identify tryptic-digested polypeptides that contained splice variant antigens in 63 primary breast and ovarian cancer samples. This method found, on average, 1.7 predicted splice variant antigens per sample, with up to 30% more alternative splicing events in tumours than in normal tissues. Notably, Kahles et al.77 also reported several known (SF3B1 and U2AF1) and novel (transcriptional adaptor 1 (TADA1), serine–threonine protein phosphatase PPP2R1A and isocitrate dehydrogenase 1 (IDH1)) splicing quantitative trait loci that were associated with alternative splicing events in 385 genes, suggesting that these loci are important for predicting the burden of splice variant antigens.
While these studies have demonstrated TSAs derived from cancer-specific splice junctions, further work will be needed to refine the computational methods for splice variant antigen prediction. Particular emphasis is needed on identifying novel splice junctions that are likely to yield mRNA isoforms that will not undergo nonsense-mediated decay95. To address this problem, improved full-length mRNA isoform inference procedures or hybrid (that is, long- and short-read) RNA-seq algorithms will need to be developed. These procedures would identify the full-length splice variant transcript, allowing for filtering of transcripts that do not contain premature stop codons that could subsequently trigger nonsense-mediated decay.
While tumour-specific splice variants of particular genes have been described in multiple tumour types, there are currently no reports of the use of splice variant antigens in personalized therapies. For example, the presence of tumour-associated splice variants has been described in select genes, including receptor for hyaluronan-mediated motility (RHAMM; two tumour-enriched variants, RHAMM-48 and RHAMM-147 in multiple myeloma)96 and Wilms tumour protein 1 (WT1; one variant, E5+, enriched in multiple cancers)97,98,99. WT1-derived peptides have been studied as a therapeutic target in leukaemias100,101,102,103,104 and in lung105 and kidney cancers106; however, these trials did not use epitopes specific for the E5+ splice variant. Additionally, an HLA–B44-restricted epitope derived from a variant of the minor histocompatibility antigen HMSD (HMSD-v) selectively expressed by primary haematological malignant cells (including those of myeloid lineage as well as multiple myeloma), but also by normal mature dendritic cells, was observed to be targeted by the CD8+ cytotoxic T cell clone 2A12-CTL107. Co-incubation of 2A12-CTL with primary AML cells conferred tumour resistance to immunodeficient mice after injection, suggesting that this HMSD-v derived antigen is a viable target for immunotherapy. Finally, Vauchy et al.108 described a CD20 splice variant (D393–CD20) whose expression is detectable in transformed B cells and upregulated in various B cell lymphomas. They subsequently demonstrated the capacity of D393–CD20-derived epitope vaccines to trigger both CD4+ and CD8+ T cell responses in HLA-humanized transgenic mice, supporting the use of CD20 splice variant epitopes for targeted immunotherapies in B cell malignancies.
Gene fusion neoantigens
Gene fusion occurrence in cancer
Gene fusions were originally identified in leukaemia109, with subsequent observations in bladder110, breast111, renal112, colon113 and lung cancers114 (among others). Similar to splice variants, gene fusion proteins have been a focus of study in leukaemia (particularly AML, acute lymphocytic leukaemia (ALL), and chronic myeloid leukaemia (CML)115) but also sarcomas116, where SNV burden is limited. These cancers contain conserved gene fusions, some of which are observed in nearly 100% of cancer subtypes (for example, t(11;22)(p13;q12) in synovial sarcoma117). Because gene fusions are often driver mutations of certain tumours, compounds aimed at inhibiting fusion protein function have been clinically successful118. Immunotherapies directed against driver mutation gene fusions may be especially beneficial, as they would directly target the source of oncogenesis. However, while driver mutation expression has been demonstrated to be highly clonal in early cancers119, studies in non-small-cell lung cancer have demonstrated highly heterogeneous driver alterations119, frequent loss of HLA heterozygosity120 and epigenetic silencing of neoantigen-containing genes occurring in later disease121, all of which may contribute to immune escape. As such, targeting of a single driver mutation may limit long-term therapeutic efficacy, whereby therapy-resistant sub-clones with differential driver mutations and HLA expression profiles may arise. Although overall gene fusion frequency is relatively low compared to SNV and INDEL mutations, they can be shared within and between different tumour types122, making them identifiable through targeted methods (for example, fluorescence in situ hybridization) and potentially targetable by universal (as opposed to patient-specific) neoantigen-based strategies.
Prediction tools for gene fusion neoantigens
Using current genomic techniques, gene fusions are typically identified through the alignment of fusion-containing reads from RNA-seq to more than one reference gene. In addition to general gene fusion callers123, several personalized gene fusion neoantigen-calling pipelines have been developed, including INTEGRATE-neo, which is specifically designed for the prediction of gene fusion neoantigens124. Using INTEGRATE-neo for analysis of the TCGA prostate adenocarcinoma cohort, 1,761 gene fusions were identified in 333 patient samples that generated 2,707 fusion transcript isoforms. Among this set, 61 (3.5% of the total) gene fusions were identified in >1 patient. Furthermore, 1,600 fusion junction peptides were identified from the 2,707 transcripts, of which 240 (15%) were predicted HLA binders124. Notably, the binding affinity scores for these 240 predicted neoantigens were skewed toward tighter affinity, suggesting that predicted fusion-derived neoantigens might have substantially better MHC binding capacity than SNV neoantigens. In addition to INTEGRATE-neo, several other tools have been described for gene fusion neoantigen calling, including pVACfuse (which performs neoantigen epitope calling using fusion variants reported from INTEGRATE-neo), NeoepitopePred125, Antigen.garnish47 and Epidisco46.
Clinical studies with gene fusion neoantigens
Clinical trials targeting gene fusion neoantigens have been pursued in CML (targeting the BCR–ABL fusion) and paediatric sarcomas. Pinilla-Ibarz et al.126 demonstrated that three of six patients with CML receiving a high dose of a BCR–ABL fusion protein breakpoint peptide vaccine developed antigen-specific T cell responses, although no cytotoxic response was observed. Although this phase I study was designed to assess safety and not clinical efficacy, one patient demonstrated transient loss of BCR–ABL mRNA, one patient experienced transient and partial cytogenic response during vaccination, and two patients progressed to an accelerated phase of disease during the study period. A follow-up phase II trial from the same group similarly demonstrated evidence of vaccine safety and measurable immunogenic response, but no evidence of clinical efficacy127. Another trial, summarized in a publication from Mackall et al.128, studied the effects of dendritic cells pulsed with tumour-specific translocation breakpoints and E7, a peptide known to bind to HLA-A2 (given alongside autologous T cells +/- IL-2 and, serving as a control, influenza vaccinations) in patients with Ewing sarcoma and alveolar rhabdomyosarcoma. Compared with the 31% five-year overall survival in patients who underwent control apheresis, immunotherapy-treated patients had 43% five-year overall survival with minimal toxicity. These studies (among others129,130) underscore the potential for gene fusion neoantigens as universal off-the-shelf therapeutics, although their current clinical efficacy remains modest. This may be related in part to therapies only targeting a single gene fusion epitope, allowing for resistant sub-clones to arise in the later disease course119,120. While an off-the-shelf approach has clear logistical merit, the identification and application of multiple patient-specific gene fusion epitopes may improve therapeutic efficacy.
Currently, few studies have applied patient-specific fusion proteins predicted through DNA- and/or RNA-seq methods for therapeutic vaccination. One recent example from Yang et al.131 demonstrated the capacity of INTEGRATE-neo-derived fusion epitopes from head and neck cancers, including fusion epitopes derived from cancers with low overall mutational burden, to generate ex vivo activation of host and healthy donor T cells. Large-cohort clinical studies (for example, PGV-001 (refs46,94)) are currently underway that will include gene fusion neoantigens among the set of targeted TSAs. Future use of this potential class of neoantigens, alone or in combination, will require larger clinical trials with more robust clinical and immunological endpoints.
Endogenous retroelement antigens
Retrotransposons in cancer
Retrotransposons are mobile genetic elements capable of self-replication through transcription and reverse transcription from genomic DNA132. They can be broadly divided into long-terminal repeat (LTR, also known as retroviral-like) and non-LTR subclasses, which differ in their genomic structures and replication mechanisms132. Retrotransposons can be expressed in cancer through epigenetic dysregulation, either through inherently low methylation states11,133,134 or following pharmacological induction of demethylation135,136,137,138, resulting in transcription (and potential translation) of retroviral TSAs139. Among the many classes of retrotransposons, long interspersed nuclear elements (LINEs, a class of non-LTR retrotransposon) have been best characterized in terms of their ability to impact cancer biology. LINE-1 has been shown to induce cancer cell apoptosis140, trigger adenomatous polyposis coli (APC)-mediated tumorigenesis in colon cancer141 and associate with clinical features and changes in cellular morphology in breast cancer142,143, among other roles.
Endogenous retroviruses (ERVs), a type of LTR retrotransposon in mammals, are remnants of exogenous retroviruses that have been incorporated into the genome throughout evolution144. Human ERVs (hERVs) impact the pathogenesis and progression of cancers, including melanoma, lymphoma, leukaemia and ovarian, prostate, urothelial and renal carcinomas134,145,146,147,148,149,150,151,152,153. Transcription of tumour-specific or enriched hERVs arises through epigenetic dysregulation of the cancer genome (which can be either inherent to the epigenetic state of the cancer or pharmacologically induced through epigenetic modulating agents), resulting in the expression of hERV-containing genomic regions otherwise not observed under physiological conditions136,138. These tumour-specific or enriched hERVs can impact both the innate and adaptive immune system through distinct mechanisms. With the innate immune system, hERVs signal through innate sensors, most commonly the RIG-I-like pathway recognition of viral double-stranded RNAs136,138. This results in downstream nuclear factor-κB (NF-κB)-mediated inflammation, with release of type I interferon, which causes immune activation and the expression of class I MHCs on tumour cells. Additionally, hERV-derived protein antigens can induce B cell and T cell activation154,155,156. Therefore, it has been proposed that tumour-specific hERV antigens could be applied to antitumour adoptive cellular therapies and therapeutic vaccines.
In addition to INDEL-derived neoantigens, hERVs have been proposed as key drivers of antitumour immunity in ccRCC11,157. In ccRCC, hERVs demonstrate baseline expression in the tumour without exogenous pharmacological epigenetic modulation, with expression of these hERVs showing strong association with both clinical prognosis and response to immunotherapy11,157. A 2015 study from Rooney et al.158 provided an initial genomic evaluation into the interaction between hERVs and the tumour immune microenvironment, demonstrating three of 66 hERVs (ERVH-5, ERVH48-1, ERVE-4; identified in a previous study from Mayer et al.159) to have tumour-specific expression and correlate with a cytotoxicity signature (granzyme A and perforin-1) in several cancers. On the basis of this study as well as several other translational studies showing the presence of an hERV-specific T cell response in ccRCC155,160, our group performed comprehensive analyses into the role of hERVs in ccRCC11,157. From immunogenomic analysis of hERVs in ccRCC, we demonstrated hERV-derived signatures to be the best predictor of patient prognosis, outperforming both clinical stage and M1–M4 molecular subtyping11. Additionally, the expression of tumour-specific hERV 4700 in pretreatment ccRCC samples was strongly associated with post-treatment response rates to anti-programmed cell death 1 (PD1) therapy. As such, hERV-derived antigens may be a viable alternative TSA target in ccRCC. Additionally, recent evidence suggests a potential role for hERVs in the modulation of low-grade glioma (where SNV burden is among the lowest of any cancer)11 and testicular cancer (particularly those with KIT mutations), in which global DNA hypomethylation is associated with high hERV expression133.
Computational methods for the quantification of retroelement expression
Several computational methods for retroelement quantification currently exist, with the majority providing quantification of ERV-like or retrotransposon-like elements (partial or full-length) rather than full-length, intact ERVs at specific genomic coordinates. This is due to the historic lack of well-annotated ERV references containing full proviral sequences and coordinates (rather than segments of ERV-like elements), which have only recently been published, to allow for mapping of full-length, intact ERVs161,162. The most well-known tool is RepeatMasker, designed to identify interspersed repeats and low-complexity sequences of any class, including simple and tandem repeats, segmental duplications, and interspersed repeats (including ERV-like elements, LINEs and short interspersed nuclear elements (SINEs), LTRs and other classes)163. RepeatMasker used in its default state is not optimal for the detection of ERVs. However, many ERV-specific databases (for example, HERVd164, HESAS165 and EnHERV166) have subsequently been generated using RepeatMasker. A more recent quantifier designed by our group, aimed specifically for the analysis of hERVs from RNA-seq data, is hervQuant11, which quantifies full-length, intact hERV proviral sequences. The hervQuant reference is derived from Vargiu et al.161, which compiled genomic coordinates for 3,173 full-length hERV proviruses. Notably, hervQuant provided the first description of a broad genomic screening method for tumour-specific hERV antigens.
Because no tools are currently available to identify retroelement TSAs, the retroelement or ERV quantifiers described above must be paired with downstream epitope prediction software (for example, NetMHCpan27) for retroelement antigen binding predictions. Additionally, because retroelements are present in the genome of both tumour and normal tissues, the prediction of tumour-specific retroelements provides unique challenges. Unlike the identification of neoantigens, retroelement TSAs must be derived through differential expression analysis of tumour versus normal-tissue RNA-seq. While hERVs and other retroelements share common homology among their overall sequences, which might theoretically make them unsuitable targets for TSA therapeutic approaches, they also exhibit highly unique regions specific to each hERV, capable of generating equally unique peptide epitopes11. Our analysis of hERV homology during the design of hervQuant revealed that only a minority of hERVs contain >95% sequence homology with one or more other hERVs, providing the basis for our ability to differentiate hERVs from short-read RNA-seq data. Such hERV unique regions can be leveraged for hERV-based TSA therapies, as long as one can confirm the specificity of expression of that particular hERV within a tumour. Additionally, evidence of hERV-specific T cells found natively within the tumour immune microenvironment (for example, hERV 4700 (ref.11) and CT RCC HERV-E167) suggests a lack of thymic central tolerance against these hERV-specific epitopes.
Translational relevance of tumour-specific hERV targets
Several studies have described the translational application of tumour-specific hERV targets. A 2016 study from Cherkasova et al.155 identified a CD8+ T cell clone from a patient with regressing ccRCC and found the clone to have tumour-specific cytotoxicity in vitro. The CTL recognized an antigen from a specific hERV, CT RCC HERV-E — which was the same as one of the tumour-specific hERVs (ERVE-4) described by Rooney et al.158 and was also identified during our screen of differentially expressed hERVs in ccRCC (hERV 2256). This particular CTL clone is being studied in clinical trials for adoptive T cell therapy in metastatic ccRCC167. Our analysis also identified a second hERV (hERV 4700) with preferential expression in ccRCC compared to normal tissues, evidence of translation, and the presence of tumour-infiltrating CTLs specific for hERV-4700 gag- and pol-derived antigens of the virus11.
Other alternative TSAs
HLA somatic mutation-derived neoantigens
Several studies have described somatic mutations in tumour HLAs that allow for altered T cell recognition. This was first described by Brandle et al.168, where mutated HLA-A2*0201 in a ccRCC tumour promoted an antitumour T cell response. This study did not elucidate whether the mechanism for T cell response was a result of TCR recognition of the antigen presented by the mutated HLA or whether the recognition was against the HLA molecule itself. Huang et al.169 later demonstrated a similar finding in metastatic melanoma, with evidence that tumour-specific T cells may recognize an unknown antigen or set of antigens presented on mutated tumour HLA-A11. Together, these studies provided early evidence for potential targeting of novel antigens with specificity of binding to somatically mutated HLA on the tumour. A recent publication from Shulka et al.22 presented whole-exome-based HLA-typing software, POLYSOLVER, that is able to call HLA somatic mutations with high prediction power, validated by RNA-seq (estimated sensitivity 94.1%, specificity 53.3%). More recently, HLAProfiler improved upon the breadth and accuracy of HLA somatic mutation calls and is able to work from RNA-seq data alone26. Combined with existing tools capable of predicting antigen binding directly from HLA sequences (for example, NetMHCpan), it is possible to predict for sets of antigens with specificity for the mutated tumour HLA, and thus specificity for antitumour T cell responses. Notably, a more advanced version of NetMHCpan is theoretically able to predict MHC binding to novel MHC molecules (including those containing mutations) through machine-learning prediction of MHC binding based on the amino acid sequence of the MHC variant27.
Post-translational TSAs
TSAs can arise post-translationally in tumours, with the potential to be targets for therapy, but they are difficult to predict with current computational tools. Post-translational splicing may occur in human cancers, resulting in the excision of a polypeptide segment followed by subsequent ligation of the free carboxyl-terminal with the amino-terminal of a new peptide74,75. Additionally, a second class of antigens, known as T cell epitopes associated with impaired peptide processing (TEIPP), has been described as being presented on transporter involved in antigen processing (TAP)-deficient, MHC-low tumours and as being recognized by a TEIPP-specific T cell population170,171,172,173,174. Interestingly, these epitopes are non-mutated and derived from housekeeping genes. Yet they are not presented on normal cells. TEIPP-specific T cells can escape thymic selection in wild-type mice (but not in TAP1-deficient mice), making them promising candidates for antitumour therapeutic targets.
Challenges and future directions
Among the challenges impeding broad clinical application of alternative TSAs as therapeutics is the need to increase the sensitivity and accuracy of epitope prediction. The computational methods described above have provided avenues to predict a greater number of TSAs from a broader variety of genomic sources. However, methods both upstream and downstream of these algorithms can generally be applied to improve the prediction performance for all TSA classes. Here, we highlight several strategies that may universally increase the number and accuracy of all TSA predictions: improvement of MHC epitope binding predictions, algorithms for the direct prediction of TSA generation and immunogenicity, and mass spectrometry approaches to improve TSA calling accuracy.
MHC epitope calling
Most TSA therapeutic vaccine studies to date have focused on the use of predicted MHC I binding epitopes, largely due to the classical hypothesis that CD8+ T cells play a greater role in antitumour immunity than do CD4+ T cells, as well as the better performance of MHC I epitope prediction algorithms than of MHC II epitope predictors. Despite this, further improvements will be required for both MHC I and II prediction algorithms to identify greater numbers of accurately predicted TSAs. A recent analysis from our group demonstrated that the accuracy of MHC I binding affinity predictions by NetMHCpan varied greatly by allele type, with performance measures being strongly correlated with the proportion of training data epitopes that were ‘binders’ (KD ≤ 500 nM, the generally accepted cut-off within the field for MHC binding), and less so with the amount of total training data per allele175. As such, alleles with fewer ‘binders’ in the training set suffered from poor sensitivity and specificity, suggesting that more high-quality data will be necessary for the application of MHC I predictors for clinical TSA prediction.
Regarding MHC II predictions, recent preclinical and clinical studies have suggested the importance of MHC II binding neoantigens in promoting antitumour immunity. A study from Kreiter et al.176 was the first to describe MHC I-predicted neoantigens in fact being presented on MHC II, subsequently triggering CD4+ T cell responses. The relevance of SNV-specific CD4+ T cells in antitumour immunity is further supported by an earlier study from Tran et al.177, whereby infusion of an ERBB2 interacting protein (ERBB2IP) mutation-specific CD4+ T cell population abrogated tumour growth for 35 months in a patient with metastatic cholangiocarcinoma. Clinical studies from Sahin et al.2,3 confirmed the importance of CD4+ T cell responses in human trials, providing evidence in support of the clinical importance of MHC II TSAs. Despite this evidence, a major hurdle faced by computational prediction methods for MHC II epitopes arises from the open-binding cleft structure of the MHC II complex. This structure results in relatively promiscuous binding of epitopes compared with MHC I, whereby the binding core of the longer class II epitope must be accurately predicted before binding affinity can subsequently be calculated178.
Recent improvements have been made in the computational prediction of MHC II epitope binding affinity, largely facilitated by the application of machine-learning algorithms trained on large, validated epitope datasets. Many earlier algorithms focused on the identification of the epitope binding core, with predictions based on the interactions between this peptide core and the MHC complex. Neilson and colleagues179 first described NN-align, which provided MHC II binding predictions trained on both the peptide-core and flanking-region characteristics, which significantly improved MHC II binding prediction performance. Although the binding affinity of an epitope is primarily determined by its peptide core, flanking-region characteristics can also influence the binding affinity. NN-align was later adapted as the core algorithm for NetMHCIIpan by Andreatta et al.180, which further improved performance, and led to the description of alternative epitope–MHC interactions. Even with these improvements to MHC II binding prediction, the performance characteristics of state-of-the-art algorithms still lag behind MHC I binding predictors. While the importance of MHC II epitopes in promoting antitumour immunity has primarily been observed with SNV neoantigens, it is expected that alternative TSAs would similarly be applicable as MHC II epitopes. As such, increasing both the breadth of available TSAs from alternative sources and the improvements to MHC II epitope prediction can together provide a concerted strategy to multiplicatively increase the targetable pool of tumour antigens.
Direct prediction of TSA generation and immunogenicity
In addition to MHC binding affinity prediction, new methods for direct prediction of TSA generation and immunogenicity might aid in the clinical selection of therapeutic epitopes. The majority of neoantigen prediction algorithms currently rely on predicted peptide–MHC binding affinity as the primary method for epitope screening. However, preclinical and clinical studies have demonstrated that only a minority of all neoantigen candidates are capable of producing immune responses2,3,176,181. One such explanation for this high false-positive rate is that the current binding prediction tools do not account for other steps involved in MHC peptide processing182. A study from Pearson et al.183 demonstrated that MHC class I associated peptides (MAPs; that is, epitopes) were derived from only 10% of exomic sequences expressed in B lymphocytes, with 41% of protein-coding genes generating no MAPs. Using features of transcripts and proteins associated with efficient MAP production, they generated a logistic regression model to predict whether a gene is capable of MAP generation.
Another approach to improving TSA prediction is to directly predict epitope immunogenicity. As we briefly mentioned above, Neopepsee is a new tool that incorporates a machine-learning algorithm trained on HLA alleles that generate T cell responses to directly predict the immunogenicity of putative neoantigens44. Compared with conventional binding affinity metrics, Neopepsee predicted immunogenicity in two external validation datasets with significantly improved sensitivity and specificity, providing evidence in support of direct immunogenicity prediction approaches. Because the Neopepsee algorithm was trained on a broad set of HLA alleles rather than specifically using TSA epitope immunogenicity, biological differences between self-derived neoantigens and the non-self epitopes of the training set may be a limiting factor for algorithmic performance. With an increasingly growing number of clinical trials collecting neoantigen immunogenicity data, future algorithms trained specifically on TSAs may potentially provide even better predictive capabilities.
Mass spectrometry approaches
Apart from computational TSA prediction, mass spectrometry-based peptidomic approaches have been applied for the identification of tumour antigens184. The identification of endogenously presented epitopes by mass spectrometry began in the early 1990s185,186. The first peptide antigens were discovered with manual interpretation of tandem mass spectrometry; however, computational methods are now the routine strategy to make comparisons between tandem mass spectrometry and peptide sequences on proteomics databases. While conceptually similar to genomic alignment and sequencing, tandem mass spectrometry sequencing is substantially more error prone and less sensitive. Standard proteomics experiments with complex peptide mixtures from well characterized biological samples are typically only able to identify ~25% of the tandem mass spectra in a proteomics database187,188. In addition to the computational difficulties with sequence identification, peptides can undergo rearrangements in the mass spectrometer, generating sequences that were not present in the original biological sample189,190. Despite these challenges, progress has been made in confirming predicted neoantigens using mass spectrometry. Immunogenomic methods have been used to generate virtual peptidomes from tumour sequencing data, and neoantigens have been identified191,192. A recent study by Laumont et al.193 used mass spectrometry approaches and observed that approximately 90% of the identified TSAs from two mouse cancer cell lines and seven human primary tumours were derived from noncoding regions, including introns, alternative reading frames, noncoding exons, untranslated region–exon junctions, structural variants and endogenous retroelements. Notably, these noncoding regions are not identified through current exome or transcriptome-based sequencing approaches. This study underscores the potential importance of the alternative TSAs and provides strong evidence for their application in therapy design, importantly demonstrating that classical SNV neoantigens may comprise only a minority of the total TSA repertoire. While these studies have enabled neoantigen discovery on a large scale, the limitations of tandem mass spectrometry alignment and the possibility of unexpected peptide rearrangements mean that suspected neoantigens should be confirmed by direct comparison of the sample’s tandem mass spectrum with that of the synthetic peptide175,191,194.
Conclusion
Conventional SNV neoantigens remain the most well-studied class of TSA, with distinct advantages in terms of ease of prediction, prevalence in a wide cohort of patients and promising preclinical and clinical therapeutic evidence of immunogenicity. While SNV neoantigens will continue to be a driving force for therapeutic vaccine development in the coming years, many groups have broadened the search for other alternative TSAs derived from self- and non-self-antigens. While certain sources of alternative TSAs have been studied for decades (for example, gene fusion proteins and viral antigens), the advent of powerful computational methods for the patient-specific prediction of TSAs has expanded the breadth of targets available for potential clinical applications. Unlike SNV neoantigens, which are largely patient specific in expression12, some classes of alternative TSAs are shared among the population (for example, splice variant antigens, gene fusion antigens and hERV antigens), making them ideal for off-the-shelf therapies. Additionally, many of these peptide sequences are highly dissimilar from germline sequences (for example, frameshifts), allowing for potentially greater immunogenicity than SNV neoantigens. Thus, alternative TSAs should play a major role in the future of cancer immunotherapy.
References
Yarchoan, M. et al. Targeting neoantigens to augment antitumour immunity. Nat. Rev. Cancer 17, 209–222 (2017).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Gubin, M. M. et al. Tumor neoantigens: building a framework for personalized cancer immunotherapy. J. Clin. Invest. 125, 3413–3421 (2015).
Hacohen, N. et al. Getting personal with neoantigen-based therapeutic cancer vaccines. Cancer Immunol. Res. 1, 11–15 (2013).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Keskin, D. B. et al. Neoantigen vaccine generates intratumoral T cell responses in phase Ib glioblastoma trial. Nature 565, 234–239 (2019).
Hilf, N. et al. Actively personalized vaccination trial for newly diagnosed glioblastoma. Nature 565, 240–245 (2019).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Smith, C. C. et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J. Clin. Invest. 128, 4804–4820 (2018).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830 (2018).
Mertens, F., Antonescu, C. R. & Mitelman, F. Gene fusions in soft tissue tumors: recurrent and overlapping pathogenetic themes. Genes Chromosomes Cancer 55, 291–310 (2016).
Wang, Y. et al. Recurrent fusion genes in leukemia: an attractive target for diagnosis and treatment. Curr. Genomics 18, 378–384 (2017).
Pellagatti, A. et al. Impact of spliceosome mutations on RNA splicing in myelodysplasia: dysregulated genes/pathways and clinical associations. Blood 132, 1225–1240 (2018).
Bartel, F., Taubert, H. & Harris, L. C. Alternative and aberrant splicing of MDM2 mRNA in human cancer. Cancer Cell 2, 9–15 (2002).
Perz, J. F. et al. The contributions of hepatitis B virus and hepatitis C virus infections to cirrhosis and primary liver cancer worldwide. J. Hepatol. 45, 529–538 (2006).
Ambrosio, M. R. & Leoncini, L. in Tropical Hemato-Oncology (eds Droz, J.-P. et al.) 127–141 (Springer International Publishing, 2015).
Mahieux, R. & Gessain, A. HTLV-1 and associated adult T cell leukemia/lymphoma. Rev. Clin. Exp. Hematol. 7, 336–361 (2003).
Mesri, E. A., Cesarman, E. & Boshoff, C. Kaposi’s sarcoma herpesvirus/ Human herpesvirus-8 (KSHV/HHV8), and the oncogenesis of Kaposi’s sarcoma. Nat. Rev. Cancer 10, 707–719 (2010).
Harrington, W. J., Wood, C. & Wood, C. in DNA Tumor Viruses (eds Pipas, J. & Damania, B.) 683–702 (Springer, 2009).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class i HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Szolek, A. et al. OptiType: precision HLA typing from next-generation sequencing data. Bioinformatics 30, 3310–3316 (2014).
Bai, Y., Wang, D. & Fury, W. PHLAT: inference of high-resolution HLA types from RNA and whole exome sequencing. Methods Mol. Biol. 1802, 193–201 (2018).
Ka, S. et al. HLAscan: genotyping of the HLA region using next-generation sequencing data. BMC Bioinformatics 18, 258 (2017).
Buchkovich, M. L. et al. HLAProfiler utilizes k-mer profiles to improve HLA calling accuracy for rare and common alleles in RNA-seq data. Genome Med. 9, 86 (2017).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide–MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Rajasagi, M. et al. Systematic identification of personal tumor-specific neoantigens in chronic lymphocytic leukemia. Blood 124, 453–462 (2014).
Soria-Guerra, R. E. et al. An overview of bioinformatics tools for epitope prediction: implications on vaccine development. J. Biomed. Inform. 53, 405–414 (2015).
Zhang, Q. et al. Immune epitope database analysis resource (IEDB-AR). Nucleic Acids Res. 36, W513–W518 (2008).
Linnebacher, M. et al. Frameshift peptide-derived T cell epitopes: a source of novel tumor-specific antigens. Int. J. Cancer 93, 6–11 (2001).
Thibodeau, S. N., Bren, G. & Schaid, D. Microsatellite instability in cancer of the proximal colon. Science 260, 816–819 (1993).
Ionov, Y. et al. Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature 363, 558–561 (1993).
Sakurada, K. et al. RIZ, the retinoblastoma protein interacting zinc finger gene, is mutated in genetically unstable cancers of the pancreas, stomach, and colorectum. Genes Chromosomes Cancer 30, 207–211 (2001).
De Smedt, L. et al. Microsatellite instable versus stable colon carcinomas: analysis of tumour heterogeneity, inflammation and angiogenesis. Br. J. Cancer 113, 500–509 (2015).
Dolcetti, R. et al. High prevalence of activated intraepithelial cytotoxic T lymphocytes and increased neoplastic cell apoptosis in colorectal carcinomas with microsatellite instability. Am. J. Pathol. 154, 1805–1813 (1999).
Maby, P. et al. Correlation between density of CD8+ T cell infiltrate in microsatellite unstable colorectal cancers and frameshift mutations: a rationale for personalized immunotherapy. Cancer Res. 75, 3446–3455 (2015).
Tougeron, D. et al. Tumor-infiltrating lymphocytes in colorectal cancers with microsatellite instability are correlated with the number and spectrum of frameshift mutations. Mod. Pathol. 22, 1186–1195 (2009).
Saeterdal, I. et al. TGF betaRII frameshift-mutation-derived CTL epitope recognised by HLA-A2-restricted CD8+ T cells. Cancer Immunol. Immunother. 50, 469–476 (2001).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Gong, J. et al. Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: a comprehensive review of registration trials and future considerations. J. Immunother. Cancer 6, 8 (2018).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Hundal, J. et al. pVAC-Seq: a genome-guided in silico approach to identifying tumor neoantigens. Genome Med. 8, 11 (2016).
Kim, S. et al. Neopepsee: accurate genome-level prediction of neoantigens by harnessing sequence and amino acid immunogenicity information. Ann. Oncol. 29, 1030–1036 (2018).
Bjerregaard, A. M. et al. MuPeXI: prediction of neo-epitopes from tumor sequencing data. Cancer Immunol. Immunother. 66, 1123–1130 (2017).
Rubinsteyn, A. et al. Computational pipeline for the PGV-001 neoantigen vaccine trial. Front. Immunol. 8, 1807 (2018).
Rech, A. J. et al. Tumor immunity and survival as a function of alternative neopeptides in human cancer. Cancer Immunol. Res. 6, 276–287 (2018).
Zhou, Z. et al. TSNAD: an integrated software for cancer somatic mutation and tumour-specific neoantigen detection. R. Soc. Open Sci. 4, 170050 (2017).
Saunders, C. T. et al. Strelka: Accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Saeterdal, I. et al. Frameshift-mutation-derived peptides as tumor-specific antigens in inherited and spontaneous colorectal cancer. Proc. Natl Acad. Sci. USA 98, 13255–13260 (2001).
Inderberg, E. M. et al. T cell therapy targeting a public neoantigen in microsatellite instable colon cancer reduces in vivo tumor growth. Oncoimmunology 6, e1302631 (2017).
Jayasinghe, R. G. et al. Systematic analysis of splice-site-creating mutations in cancer. Cell Rep. 23, 270–281 (2018).
Yang, Y. et al. Aberrant splicing induced by missense mutations in BRCA1: clues from a humanized mouse model. Hum. Mol. Genet. 12, 2121–2131 (2003).
Nyström-Lahti, M. et al. Missense and nonsense mutations in codon 659 of MLHI cause aberrant splicing of messenger RNA in HNPCC kindreds. Genes Chromosomes Cancer 26, 372–375 (1999).
Zhang, K. et al. Patterns of missplicing caused by RB1 gene mutations in patients with retinoblastoma and association with phenotypic expression. Hum. Mutat. 29, 475–484 (2008).
Wadt, K. et al. A cryptic BAP1 splice mutation in a family with uveal and cutaneous melanoma, and paraganglioma. Pigment Cell Melanoma Res. 25, 815–818 (2012).
Chen, L. L. et al. A mutation-created novel intra-exonic pre-mRNA splice site causes constitutive activation of KIT in human gastrointestinal stromal tumors. Oncogene 24, 4271–4280 (2005).
Smart, A. C. et al. Intron retention is a source of neoepitopes in cancer. Nat. Biotechnol. 36, 1056–1058 (2018).
Jung, H. et al. Intron retention is a widespread mechanism of tumor-suppressor inactivation. Nat. Genet. 47, 1242–1248 (2015).
Dvinge, H. & Bradley, R. K. Widespread intron retention diversifies most cancer transcriptomes. Genome Med. 7, 45 (2015).
Kawakami, S. A. et al. The intronic region of an incompletely spliced gp100 gene transcript encodes an epitope recognized by melanoma-reactive tumor-infiltrating lymphocytes. J. Immunol. 159, 303–308 (1997).
Coulie, P. G. et al. A mutated intron sequence codes for an antigenic peptide recognized by cytolytic T lymphocytes on a human melanoma. Proc. Natl Acad. Sci. USA 92, 7976–7980 (1995).
Uenaka, A. et al. Cryptic CTL epitope on a murine sarcoma Meth A generated by exon extension as a novel mechanism. J. Immunol. 170, 4862–4868 (2003).
Boultwood, J. et al. The role of splicing factor mutations in the pathogenesis of the myelodysplastic syndromes. Adv. Biol. Regul. 54, 153–161 (2014).
Yip, B. H. et al. Impact of splicing factor mutations on pre-mRNA splicing in the myelodysplastic syndromes. Curr. Pharm. Des. 22, 2333–2344 (2016).
Weiss, R. B. et al. Slippery runs, shifty stops, backward steps, and forward hops: -2, -1, +1, +2, +5, and +6 ribosomal frameshifting. Cold Spring Harb. Symp. Quant. Biol. 52, 687–693 (1987).
Saulquin, X. et al. +1 Frameshifting as a novel mechanism to generate a cryptic cytotoxic T lymphocyte epitope derived from human interleukin 10. J. Exp. Med. 195, 353–358 (2002).
Macejak, D. G. & Sarnow, P. Internal initiation of translation mediated by the 5’ leader of a cellular mRNA. Nature 353, 90–94 (1991).
Bullock, T. N. J. et al. Initiation codon scanthrough versus termination codon readthrough demonstrates strong potential for major histocompatibility complex class I–restricted cryptic epitope expression. J. Exp. Med. 186, 1051–1058 (1997).
Bullock, T. N. Ribosomal scanning past the primary initiation codon as a mechanism for expression of CTL epitopes encoded in alternative reading frames. J. Exp. Med. 184, 1319–1329 (1996).
Malarkannan, S. et al. Presentation of out-of-frame peptide/MHC class I complexes by a novel translation initiation mechanism. Immunity 10, 681–690 (1999).
Van Den Eynde, B. J. et al. A new antigen recognized by cytolytic T lymphocytes on a human kidney tumor results from reverse strand transcription. J. Exp. Med. 190, 1793–1800 (1999).
Bruce, A., Atkins, J. & Gesteland, R. tRNA anticodon replacement experiments show that ribosomal frameshifting can be caused by doublet decoding. Proc. Natl Acad. Sci. USA 83, 5062–5066 (1986).
Dalet, A. et al. An antigenic peptide produced by reverse splicing and double asparagine deamidation. Proc. Natl Acad. Sci. USA 108, E323–E331 (2011).
Hanada, K. I., Yewdell, J. W. & Yang, J. C. Immune recognition of a human renal cancer antigen through post-translational protein splicing. Nature 427, 252–256 (2004).
Liepe, J. et al. A large fraction of HLA class I ligands are proteasome-generated spliced peptides. Science 354, 354–358 (2016).
Kahles, A. et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224 (2018).
Shukla, G. C. & Singh, J. Mutations of RNA splicing factors in hematological malignancies. Cancer Lett. 409, 1–8 (2017).
Ley, T. J. et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Adamia, S. et al. A genome-wide aberrant RNA Splicing in patients with acute myeloid leukemia identifies novel potential disease markers and therapeutic targets. Clin. Cancer Res. 20, 1135–1145 (2014).
Wang, L. et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N. Engl. J. Med. 365, 2497–2506 (2011).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Kar, S. A. et al. Spliceosomal gene mutations are frequent events in the diverse mutational spectrum of chronic myelomonocytic leukemia but largely absent in juvenile myelomonocytic leukemia. Haematologica 98, 107–113 (2013).
Visconte, V. et al. Emerging roles of the spliceosomal machinery in myelodysplastic syndromes and other hematological disorders. Leukemia 26, 2447–2454 (2012).
Quesada, V. et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat. Genet. 44, 47–52 (2012).
Lee, S. C. W. et al. Modulation of splicing catalysis for therapeutic targeting of leukemia with mutations in genes encoding spliceosomal proteins. Nat. Med. 22, 672–678 (2016).
Lim, K. H. & Fairbrother, W. G. Spliceman - a computational web server that predicts sequence variations in pre-mRNA splicing. Bioinformatics 28, 1031–1032 (2012).
Mort, M. et al. MutPred Splice: machine learning-based prediction of exonic variants that disrupt splicing. Genome Biol. 15, R19 (2014).
Brooks, A. N. et al. Conservation of an RNA regulatory map between Drosophila and mammals. Genome Res. 21, 193–202 (2011).
Rogers, M. F. et al. SpliceGrapher: detecting patterns of alternative splicing from RNA-Seq data in the context of gene models and EST data. Genome Biol. 13, R4 (2012).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl Acad. Sci. USA 111, E5593–E5601 (2014).
Kahles, A. et al. SplAdder: identification, quantification and testing of alternative splicing events from RNA-Seq data. Bioinformatics 32, 1840–1847 (2016).
Denti, L. et al. ASGAL: aligning RNA-Seq data to a splicing graph to detect novel alternative splicing events. BMC Bioinformatics 19, 444 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02721043 (2019).
Shyu, A. B., Wilkinson, M. F. & Van Hoof, A. Messenger RNA regulation: to translate or to degrade. EMBO J. 27, 471–481 (2008).
Crainie, M. et al. Overexpression of the receptor for hyaluronan-mediated motility (RHAMM) characterizes the malignant clone in multiple myeloma: identification of three distinct RHAMM variants. Blood 93, 1684–1696 (1999).
Busse, A. et al. Wilms’ tumor gene 1 (WT1) expression in subtypes of acute lymphoblastic leukemia (ALL) of adults and impact on clinical outcome. Ann. Hematol. 88, 1199–1205 (2009).
Kramarzova, K. et al. Real-time PCR quantification of major Wilms tumor gene 1 (WT1) isoforms in acute myeloid leukemia, their characteristic expression patterns and possible functional consequences. Leukemia 26, 2086–2095 (2012).
Siehl, J. M. et al. Expression of Wilms’ tumor gene 1 at different stages of acute myeloid leukemia and analysis of its major splice variants. Ann. Hematol. 83, 745–750 (2004).
Mailänder, V. et al. Complete remission in a patient with recurrent acute myeloid leukemia induced by vaccination with WT1 peptide in the absence of hematological or renal toxicity. Leukemia 18, 165–166 (2004).
Kohrt, H. E. et al. Donor immunization with WT1 peptide augments antileukemic activity after MHC-matched bone marrow transplantation. Blood 118, 5319–5329 (2011).
Oka, Y. et al. Wilms tumor gene peptide-based immunotherapy for patients with overt leukemia from myelodysplastic syndrome (MDS) or MDS with myelofibrosis. Int. J. Hematol. 78, 56–61 (2003).
Rosenfeld, C., Cheever, M. A. & Gaiger, A. WT1 in acute leukemia, chronic myelogenous leukemia and myelodysplastic syndrome: therapeutic potential of WT1 targeted therapies. Leukemia 17, 1301–1312 (2003).
Chapuis, A. G. et al. Transferred WT1-reactive CD8+ T cells can mediate antileukemic activity and persist in post-transplant patients. Sci. Transl Med. 5, 174ra27 (2013).
Tsuboi, A. et al. WT1 peptide-based immunotherapy for patients with lung cancer: report of two cases. Microbiol. Immunol. 48, 175–184 (2004).
Iiyama, T. et al. WT1 (Wilms’ tumor 1) peptide immunotherapy for renal cell carcinoma. Microbiol. Immunol. 51, 519–530 (2007).
Kawase, T. et al. Alternative splicing due to an intronic SNP in HMSD generates a novel minor histocompatibility antigen. Blood 110, 1055–1063 (2007).
Vauchy, C. et al. CD20 alternative splicing isoform generates immunogenic CD4 helper T epitopes. Int. J. Cancer 137, 116–126 (2015).
Rowley, J. D. A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 243, 290–293 (1973).
Williams, S. V., Hurst, C. D. & Knowles, M. A. Oncogenic FGFR3 gene fusions in bladder cancer. Hum. Mol. Genet. 22, 795–803 (2013).
Tognon, C. et al. Expression of the ETV6-NTRK3 gene fusion as a primary event in human secretory breast carcinoma. Cancer Cell 2, 367–376 (2002).
The Cancer Genome Atlas Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
Young, L. C. et al. Identification of novel isoforms of the EML4-ALK transforming gene in non-small cell lung cancer. Cancer Res. 68, 4971–4976 (2008).
Lyu, X. et al. Detection of 22 common leukemic fusion genes using a single-step multiplex qRT-PCR-based assay. Diagn. Pathol. 12, 55 (2017).
Xiao, X. et al. Advances in chromosomal translocations and fusion genes in sarcomas and potential therapeutic applications. Cancer Treat. Rev. 63, 61–70 (2018).
Worley, B. S. et al. Antigenicity of fusion proteins from sarcoma-associated. Cancer Res. 61, 6868–6875 (2001).
Druker, B. J. et al. Efficacy and safety of a specific inhibitor of the BCR–ABL tyrosine kinase in chronic myeloid leukemia. N. Engl. J. Med. 344, 1031–1037 (2001).
Jamal-Hanjani, M. et al. Tracking the evolution of non–small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
McGranahan, N. et al. Allele-Specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271 (2017).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Yu, Y. P. et al. Identification of recurrent fusion genes across multiple cancer types. Sci. Rep. 9, 1074 (2019).
Wang, Q. et al. Application of next generation sequencing to human gene fusion detection: computational tools, features and perspectives. Brief. Bioinform. 14, 506–519 (2013).
Zhang, J., Mardis, E. R. & Maher, C. A. INTEGRATE-neo: a pipeline for personalized gene fusion neoantigen discovery. Bioinformatics 33, 555–557 (2017).
Chang, T. C. et al. The neoepitope landscape in pediatric cancers. Genome Med. 9, 78 (2017).
Pinilla-Ibarz, J. et al. Vaccination of patients with chronic myelogenous leukemia with bcr–abl oncogene breakpoint fusion peptides generates specific immune responses. Blood 95, 1781–1787 (2000).
Cathcart, K. et al. A multivalent bcr–abl fusion peptide vaccination trial in patients with chronic myeloid leukemia. Blood 103, 1037–1042 (2004).
Mackall, C. L. et al. A pilot study of consolidative immunotherapy in patients with high-risk pediatric sarcomas. Clin. Cancer Res. 14, 4850 (2008).
Bocchia, M. et al. Effect of a p210 multipeptide vaccine associated with imatinib or interferon in patients with chronic myeloid leukaemia and persistent residual disease: a multicentre observational trial. Lancet 365, 657–662 (2005).
Rojas, J. M. et al. Clinical evaluation of BCR–ABL peptide immunisation in chronic myeloid leukaemia: results of the EPIC study. Leukemia 21, 2287–2295 (2007).
Yang, W. et al. Immunogenic neoantigens derived from gene fusions stimulate T cell responses. Nat. Med. 25, 767–775 (2019).
Goodier, J. L. & Kazazian, H. H. Retrotransposons revisited: the restraint and rehabilitation of parasites. Cell 135, 23–35 (2008).
Shen, H. et al. Integrated molecular characterization of testicular germ cell tumors. Cell Rep. 23, 3392–3406 (2018).
Florl, A. R. et al. DNA methylation and expression of LINE-1 and HERV-K provirus sequences in urothelial and renal cell carcinomas. Br. J. Cancer 80, 1312–1321 (1999).
Brocks, D. et al. DNMT and HDAC inhibitors induce cryptic transcription start sites encoded in long terminal repeats. Nat. Genet. 49, 1052–1060 (2017).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563 (2018).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Jones, P. A. et al. Epigenetic therapy in immune-oncology. Nat. Rev. Cancer 19, 151–161 (2019).
Belgnaoui, S. M. et al. Human LINE-1 retrotransposon induces DNA damage and apoptosis in cancer cells. Cancer Cell Int. 6, 13 (2006).
Scott, E. C. et al. A hot L1 retrotransposon evades somatic repression and initiates human colorectal cancer. Genome Res. 26, 745–755 (2016).
Chen, L. et al. Prognostic value of LINE-1 retrotransposon expression and its subcellular localization in breast cancer. Breast Cancer Res. Treat. 136, 129–142 (2012).
Patnala, R. et al. Inhibition of LINE-1 retrotransposon-encoded reverse transcriptase modulates the expression of cell differentiation genes in breast cancer cells. Breast Cancer Res. Treat. 143, 239–253 (2014).
Löwer, R., Löwer, J. & Kurth, R. The viruses in all of us: characteristics and biological significance of human endogenous retrovirus sequences. Proc. Natl Acad. Sci. USA 93, 5177–5184 (1996).
Boller, K. et al. Human endogenous retrovirus HERV-K113 is capable of producing intact viral particles. J. Gen. Virol. 89, 567–572 (2008).
Faff, O. et al. Retrovirus-like particles from the human T47D cell line are related to mouse mammary tumour virus and are of human endogenous origin. J. Gen. Virol. 73, 1087–1097 (1992).
Wang-Johanning, F. et al. Expression of multiple human endogenous retrovirus surface envelope proteins in ovarian cancer. Int. J. Cancer 120, 81–90 (2007).
Büscher, K. et al. Expression of human endogenous retrovirus K in melanomas and melanoma cell lines. Cancer Res. 65, 4172–4180 (2005).
Wang-Johanning, F. et al. Expression of human endogenous retrovirus K envelope transcripts in human breast cancer. Clin. Cancer Res. 7, 1553–1560 (2001).
Contreras-Galindo, R. et al. Human endogenous retrovirus K (HML-2) elements in the plasma of people with lymphoma and breast cancer. J. Virol. 82, 9329–9336 (2008).
Wang-Johanning, F. et al. Detecting the expression of human endogenous retrovirus E envelope transcripts in human prostate adenocarcinoma. Cancer 98, 187–197 (2003).
Yoshida, M., Miyoshi, I. & Hinuma, Y. Isolation and characterization of retrovirus from cell lines of human adult T cell leukemia and its implication in the disease. Proc. Natl Acad. Sci. USA 79, 2031–2035 (1982).
Kalyanaraman, V. S. et al. A new subtype of human T cell leukemia virus (HTLV-II) associated with a T cell variant of hairy cell leukemia. Science 218, 571–573 (1982).
Sauter, M. et al. Human endogenous retrovirus K10: expression of Gag protein and detection of antibodies in patients with seminomas. J. Virol. 69, 414–421 (1995).
Cherkasova, E. et al. Detection of an immunogenic HERV-E envelope with selective expression in clear cell kidney cancer. Cancer Res. 76, 2177–2185 (2016).
Takahashi, Y. et al. Regression of human kidney cancer following allogeneic stem cell transplantation is associated with recognition of an HERV-E antigen by T cells. J. Clin. Invest. 118, 1099–1109 (2008).
Panda, A. et al. Endogenous retrovirus expression is associated with response to immune checkpoint pathway in clear cell renal cell carcinoma. JCI Insight 3, 121522 (2018).
Rooney, M. S. et al. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Mayer, J., Blomberg, J. & Seal, R. L. A revised nomenclature for transcribed human endogenous retroviral loci. Mob. DNA 2, 7 (2011).
Cherkasova, E. et al. Inactivation of the von Hippel–Lindau tumor suppressor leads to selective expression of a human endogenous retrovirus in kidney cancer. Oncogene 30, 4697–4706 (2011).
Vargiu, L. et al. Classification and characterization of human endogenous retroviruses; mosaic forms are common. Retrovirology 13, 7 (2016).
Tokuyama, M. et al. ERVmap analysis reveals genome-wide transcription of human endogenous retroviruses. Proc. Natl Acad. Sci. USA 115, 12565–12572 (2018).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open — 4.0. RepeatMasker http://www.repeatmasker.org/ (2013).
Paces, J. HERVd: the human endogenous retroviruses database: update. Nucleic Acids Res. 32, 50D (2004).
Kim, T. H. et al. HESAS: HERVs expression and structure analysis system. Bioinformatics 21, 1699–1700 (2005).
Tongyoo, P. et al. EnHERV: enrichment analysis of specific human endogenous retrovirus patterns and their neighboring genes. PLOS ONE 12, e0177119 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03354390 (2019).
Brandle, D. A mutated HLA-A2 molecule recognized by autologous cytotoxic T lymphocytes on a human renal cell carcinoma. J. Exp. Med. 183, 2501–2508 (1996).
Huang, J. et al. T cells associated with tumor regression recognize frameshifted products of the CDKN2A tumor suppressor gene locus and a mutated HLA class I gene product. J. Immunol. 172, 6057–6064 (2014).
Van Hall, T. et al. Selective cytotoxic T-lymphocyte targeting of tumor immune escape variants. Nat. Med. 12, 417–424 (2006).
Doorduijn, E. M. et al. TAP-independent self-peptides enhance T cell recognition of immune-escaped tumors. J. Clin. Invest. 126, 784–794 (2016).
Marijt, K. A., Doorduijn, E. M. & van Hall, T. TEIPP antigens for T cell based immunotherapy of immune-edited HLA class Ilow cancers. Mol. Immunol. https://doi.org/10.1016/j.molimm.2018.03.029 (2018).
Doorduijn, E. M. et al. T cells specific for a TAP-independent self-peptide remain naïve in tumor-bearing mice and are fully exploitable for therapy. Oncoimmunology 7, e1382793 (2018).
Marijt, K. A. et al. Identification of non-mutated neoantigens presented by TAP-deficient tumors. J. Exp. Med. 215, 2325–2337 (2018).
Lansford, J. L. et al. Computational modeling and confirmation of leukemia-associated minor histocompatibility antigens. Blood Adv. 2, 2052–2062 (2018).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Andreatta, M. et al. Machine learning reveals a non-canonical mode of peptide binding to MHC class II molecules. Immunology 152, 255–264 (2017).
Nielsen, M. & Lund, O. NN-align. An artificial neural network-based alignment algorithm for MHC class II peptide binding prediction. BMC Bioinformatics 10, 296 (2009).
Andreatta, M. et al. Accurate pan-specific prediction of peptide-MHC class II binding affinity with improved binding core identification. Immunogenetics 67, 641–650 (2015).
Saito, R. et al. Molecular subtype-specific immunocompetent models of high-grade urothelial carcinoma reveal differential neoantigen expression and response to immunotherapy. Cancer Res. 78, 3954–3968 (2018).
The problem with neoantigen prediction [editorial]. Nat. Biotechnol. 35, 97 (2017).
Pearson, H. et al. MHC class I-associated peptides derive from selective regions of the human genome. J. Clin. Invest. 126, 4690–4701 (2016).
Creech, A. L. et al. The role of mass spectrometry and proteogenomics in the advancement of HLA epitope prediction. Proteomics 18, e1700259 (2018).
Hunt, D. F. et al. Characterization of peptides bound to the class I MHC molecule HLA-A2.1 by mass spectrometry. Science 255, 1261–1263 (1992).
Falk, K. et al. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. Nature 351, 290–296 (1991).
Michalski, A., Cox, J. & Mann, M. More than 100,000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC-MS/MS. J. Proteome Res. 10, 1785–1793 (2011).
Griss, J. et al. Recognizing millions of consistently unidentified spectra across hundreds of shotgun proteomics datasets. Nat. Methods 13, 651–656 (2016).
Yaqüe, J. et al. Peptide rearrangement during quadrupole ion trap fragmentation: added complexity to MS/MS spectra. Anal. Chem. 75, 1524–1535 (2003).
Chawner, R. et al. Peptide scrambling during collision-induced dissociation is influenced by n-terminal residue basicity. J. Am. Soc. Mass Spectrom. 25, 1927–1938 (2014).
Yadav, M. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014).
Polyakova, A., Kuznetsova, K. & Moshkovskii, S. Proteogenomics meets cancer immunology: mass spectrometric discovery and analysis of neoantigens. Expert Rev. Proteomics 12, 533–541 (2015).
Laumont, C. M. et al. Noncoding regions are the main source of targetable tumor-specific antigens. Sci. Transl Med. 10, eaau5516 (2018).
van der Lee, D. I. et al. Mutated nucleophosmin 1 as immunotherapy target in acute myeloid leukemia. J. Clin. Invest. 129, 774–785 (2019).
Matsushita, H. et al. Cancer exome analysis reveals a T cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Castle, J. C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012).
Gubin, M. M. et al. Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515, 577–581 (2014).
Carreno, B. M. et al. A dendritic cell vaccine increases the breadth and diversity of melanoma neoantigen-specific T cells. Science 348, 803–808 (2015).
Gao, Q. et al. Driver fusions and their implications in the development and treatment of human cancers. Cell Rep. 23, 227–238 (2018).
Selitsky, S. R. et al. Epstein-Barr virus-positive cancers show altered B-cell clonality. mSystems 3, e00081–18 (2018).
Acknowledgements
This work was supported by the US National Institutes of Health grants F30 CA225136 (to C.C.S.), U54 CA198999 (to J.S.S.) and P50 CA058223 (to J.S.S.), as well as by a grant from the UNC University Cancer Research Fund (to B.G.V.) and a Susan G. Komen Career Catalyst Research Grant (to B.G.V.).
Author information
Authors and Affiliations
Contributions
C.C.S., S.C., S.R.S. and P.M.A. researched the data for the article. All authors provided substantial contributions to discussion of the content. C.C.S. and P.M.A. wrote the article, and C.C.S. generated figures. All authors contributed to the review and editing of the manuscript prior to submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Cancer thanks T. Van Hall, L. Delamarre and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- Apheresis
-
Medical technique used to purify various components of whole blood. Apheresis can be performed to harvest purified T lymphocytes for subsequent immunotherapeutic application.
- Artificial neural networks
-
A class of computational modelling based on biological neural networks, able to implement change based on training input and output information to form an optimized prediction model.
- Binding core
-
The segment of polypeptide on an antigenic peptide responsible for interaction with the major histocompatibility complex binding groove. The binding core is recognized as an important predictor for binding affinity, but binding is also influenced by other factors of the epitope sequence.
- Cancer–testis antigens
-
Antigens whose expression is limited to cancer cells and reproductive tissues but not adult somatic tissue.
- Cytogenic response
-
A decrease in the number of cells with a particular chromosomal trait (classically associated with the BCR–ABL gene fusion) in response to therapy.
- Doublet decoding
-
Translational process by which an amino acid is translated from a two-base-pair codon rather than the conventional three-base-pair codon. This process can result in -1 frameshifting during translation.
- Epitope
-
Specific portion of the antigen specifically recognized by a T or B cell receptor.
- HLA typing
-
Process for identifying the HLA receptor allele of a particular tissue. This can be performed through a variety of molecular or immunological techniques.
- Immunogenomic analysis
-
Study of the combined genomics of the cancer cell and immune cell components of a tumour.
- Insertion or deletion
-
(INDEL). Insertion of bases into or deletion of them from the genome of an organism, typically in the context of a mutation or genetic variation.
- KD
-
Dissociation constant that measures the concentration of a ligand necessary to reversibly bind half of its corresponding molecular pair. In the context of peptide–major histocompatibility complex (MHC) binding, this refers to the concentration of peptide necessary to bind half of all MHC molecules.
- Lynch syndrome
-
Also known as hereditary nonpolyposis colorectal cancer. An autosomal-dominant genetic disorder of DNA mismatch repair, resulting in increased risk of microsatellite instability-driven colon cancer (among other cancer types).
- Myelodysplastic syndrome
-
A class of low-grade malignancies in which abnormal bone marrow stem cells fail to fully mature.
- Negative selection
-
Process by which self-reactive T lymphocytes are deleted during T cell education in the thymus.
- Neoantigens
-
Antigens specific to the genome of the cancer cell.
- Nonsense-mediated decay
-
Checkpoint by which mRNA transcripts containing premature stop codons are eliminated in order to reduce aberrant translation.
- Post-translational splicing
-
Post-translational excision of polypeptides, with subsequent ligation of the carboxy- and amino-terminal residues.
- Predicted neoantigens
-
Genomically predicted neoantigens with unconfirmed tumour expression and/or in vivo immunogenicity.
- Quantitative trait loci
-
Sections of DNA (loci) that are correlated with particular qualitative traits (or phenotypes) in an organismal population.
- Retroelements
-
Genetic elements capable of self-amplification, found within the genome of eukaryotic organisms. Retrotransposon DNA can be transcribed into RNA, converted back into identical DNA via reverse transcription, and then inserted into the genome at particular target sites. Retroelements include retrotransposons and endogenous retroviruses.
- Retrotransposons
-
A subset of retroelement in eukaryotic cells that possesses some characteristics of retroviruses and transposes through an RNA intermediary.
- Ribosomal frameshifting
-
Process by which codons are translated in an out-of-frame manner via slippage of the ribosome into a +/- 1 or 2 base-pair position.
- Segmental duplications
-
Long segments of repeated DNA (1-400 kb) with highly conserved sequences (>90%) that exist in the genome as a result of duplication events.
- Spliceosome
-
Molecular machinery responsible for removal (splicing) of introns from pre-mRNA.
- Tandem mass spectrometry
-
Multiple-step mass spectrometry (MS), whereby the sample is first ionized for separation in the first MS stage, followed by fragmentation for separation in the second MS stage.
- Translocation breakpoints
-
Locations where two fragments of chromosome(s) are joined subsequent to chromosomal translocation.
- Tumour antigens
-
Any antigen produced by the tumour cell, typically in the setting of enriched or specific expression relative to normal tissue(s).
- Tumour-associated antigens
-
Antigens whose expression is enriched (but not specific) to cancer cells.
- Tumour-specific antigens
-
(TSAs). Antigens (molecules capable of promoting an adaptive immune response) expressed by the tumour with minimal to no expression in normal tissue.
- Viral-derived cancer antigens
-
Antigens expressed by cancer cells derived from an oncogenic viral origin.
Rights and permissions
About this article
Cite this article
Smith, C.C., Selitsky, S.R., Chai, S. et al. Alternative tumour-specific antigens. Nat Rev Cancer 19, 465–478 (2019). https://doi.org/10.1038/s41568-019-0162-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-019-0162-4
This article is cited by
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Activation of human endogenous retroviruses and its physiological consequences
Nature Reviews Molecular Cell Biology (2024)
-
Neoantigen-targeted TCR-engineered T cell immunotherapy: current advances and challenges
Biomarker Research (2023)
-
Comprehensive analysis of neoantigens derived from structural variation across whole genomes from 2528 tumors
Genome Biology (2023)
-
Overcoming on-target, off-tumour toxicity of CAR T cell therapy for solid tumours
Nature Reviews Clinical Oncology (2023)