Abstract
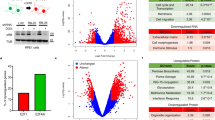
LRBA expression is induced by mitogens in lymphoid and myeloid cells. The Drosophila LRBA orthologue rugose/DAKAP550 is involved in Notch, Ras and EGFR pathways. These findings suggest that LRBA could play a role in cell types that have increased proliferative and survival capacity. Here, we show by microarray and real-time PCR analyses that LRBA is overexpressed in several different cancers relative to their normal tissue controls. We also show that LRBA promoter activity and endogenous LRBA mRNA levels are reduced by p53 and increased by E2F1, indicating that mutations in the tumor suppressors p53 and Rb could contribute to the deregulation of LRBA. Furthermore, inhibition of LRBA expression by RNA interference, or inhibition of its function by a dominant-negative mutant, leads to significant growth inhibition of cancer cells, demonstrating that deregulated expression of LRBA contributes to the altered growth properties of a cancer cell. Finally, we show that the phosphorylation of EGFR is affected by the dominant-negative mutant, suggesting LRBA plays a role in the mammalian EGFR pathway. These findings demonstrate that LRBA facilitates cancer cell growth and thus LRBA may represent a novel molecular target for cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Abbreviations
- AKAP:
-
A kinase anchor protein
- BEACH:
-
beige And CHS
- CHS1 :
-
Chediak–Higashi Syndrome 1
- CNS:
-
central nervous system
- EGFR:
-
epidermal growth factor receptor
- ER:
-
estrogen receptor
- FACS:
-
fluorescence-activated cell sorting
- FAN:
-
factor associated with neutral sphingomyelinase
- LRBA:
-
lipopolysaccharide-responsive and beige-like anchor
- LvsA:
-
large volume sphere A
- Nbea:
-
neurobeachin
- PARP:
-
poly (ADP-ribose) polymerase
- PH:
-
pleckstrin homology
- RACE:
-
rapid amplification of cDNA ends
- RB:
-
retinoblastoma protein
- rg :
-
rugose
- RNAi:
-
RNA interference
- SHIP:
-
SH2-containing inositol phosphatase
- shRNA:
-
small-hairpin RNA
- siRNA:
-
small interfering RNA
- SMART:
-
switching mechanism at 5′ end of RNA transcript
- TNF:
-
tumor necrosis factor
References
Adam-Klages S, Adam D, Wiegmann K, Struve S, Kolanus W, Schneider-Mergener J and Kronke M . (1996). Cell, 86, 937–947.
Bargonetti J and Manfredi JJ . (2002). Curr. Opin. Oncol., 14, 86–91.
Bhattacharjee A, Richards WG, Staunton J, Li C, Monti S, Vasa P, Ladd C, Beheshti J, Bueno R, Gillette M, Loda M, Weber G, Mark EJ, Lander ES, Wong W, Johnson BE, Golub TR, Sugarbaker DJ and Meyerson M . (2001). Proc. Natl. Acad. Sci. USA, 98, 13790–13795.
Blomberg N, Baraldi E, Nilges M and Saraste M . (1999). Trends Biochem. Sci., 24, 441–445.
Bloom G, Yang IV, Boulware D, Kwong KY, Coppola D, Eschrich S, Quackenbush J and Yeatman TJ . (2004). Am. J. Pathol., 164, 9–16.
Dyomin V, Chaganti S, Dyomina K, Palanisamy N, Murty V, Dalla-Favera R and Chaganti R . (2002). Genomics, 80, 158.
Han JD, Baker NE and Rubin CS . (1997). J. Biol. Chem., 272, 26611–26619.
Harborth J, Elbashir SM, Bechert K, Tuschl T and Weber K . (2001). J. Cell Sci., 114, 4557–4565.
He TC, Zhou S, da Costa LT, Yu J, Kinzler KW and Vogelstein B . (1998). Proc. Natl. Acad. Sci. USA, 95, 2509–2514.
Jogl G, Shen Y, Gebauer D, Li J, Wiegmann K, Kashkar H, Kronke M and Tong L . (2002). EMBO J., 21, 4785–4795.
Jorissen RN, Walker F, Pouliot N, Garrett TP, Ward CW and Burgess AW . (2003). Exp. Cell Res., 284, 31–53.
Kerr WG, Heller M and Herzenberg LA . (1996). Proc. Natl. Acad. Sci. USA, 93, 3947–3952.
Kwak E, Gerald N, Larochelle DA, Vithalani KK, Niswonger ML, Maready M and De Lozanne A . (1999). Mol. Biol. Cell, 10, 4429–4439.
Li D and Roberts R . (2001). Cell Mol. Life Sci., 58, 2085–2097.
Miller WR . (2002). Ann. NY Acad. Sci., 968, 37–48.
Morris C . (2002). Breast Cancer Res. Treat., 75, S51-5–S57-9 discussion.
Nagle DL, Karim MA, Woolf EA, Holmgren L, Bork P, Misumi DJ, McGrail SH, Dussault Jr BJ, Perou CM, Boissy RE, Duyk GM, Spritz RA and Moore KJ . (1996). Nat. Genet., 14, 307–311.
Ohtani K . (1999). Front Biosci., 4, D793–D804.
Segui B, Cuvillier O, Adam-Klages S, Garcia V, Malagarie-Cazenave S, Leveque S, Caspar-Bauguil S, Coudert J, Salvayre R, Kronke M and Levade T . (2001). J. Clin. Invest., 108, 143–151.
Shamloula HK, Mbogho MP, Pimentel AC, Chrzanowska-Lightowlers ZM, Hyatt V, Okano H and Venkatesh TR . (2002). Genetics, 161, 693–710.
Sherr CJ and McCormick F . (2002). Cancer Cell, 2, 103–112.
Spritz RA . (1998). J. Clin. Immunol., 18, 97–105.
Sui G, Soohoo C, Affar el B, Gay F, Shi Y and Forrester WC . (2002). Proc. Natl. Acad. Sci. USA, 99, 5515–5520.
Van Gelder RN, von Zastrow ME, Yool A, Dement WC, Barchas JD and Eberwine JH . (1990). Proc. Natl. Acad. Sci. USA, 87, 1663–1667.
Vogelstein B, Lane D and Levine AJ . (2000). Nature, 408, 307–310.
Wang JW, Howson J, Haller E and Kerr WG . (2001). J. Immunol., 166, 4586–4595.
Wang X, Herberg FW, Laue MM, Wullner C, Hu B, Petrasch-Parwez E and Kilimann MW . (2000). J. Neurosci., 20, 8551–8565.
Warrington JA, Nair A, Mahadevappa M and Tsyganskaya M . (2000). Physiol. Genomics, 2, 143–147.
Wiley HS . (2003). Exp. Cell. Res., 284, 78–88.
Acknowledgements
We thank Drs John Ninos, Steven Enkemann and Edward R Seijo for technical advice and assistance on flow cytometry analysis, microarray analysis and analytic microscopy, respectively. Various portions of this work were assisted by the Molecular Biology, Cytometry Microarray and Analytic Microscopy Core Facilities at the H Lee Moffitt Cancer Center & Research Institute. This work was supported in part by grants from the NIH RO1 DK54767, RO1 HL72523 and PO1 NS27405. WGK is The Newman Scholar of the Leukemia and Lymphoma Society.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, JW., Gamsby, J., Highfill, S. et al. Deregulated expression of LRBA facilitates cancer cell growth. Oncogene 23, 4089–4097 (2004). https://doi.org/10.1038/sj.onc.1207567
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207567
Keywords
This article is cited by
-
Phosphorylated ATF1 at Thr184 promotes metastasis and regulates MMP2 expression in gastric cancer
Journal of Translational Medicine (2022)
-
Epigenetic aging of the demographically non-aging naked mole-rat
Nature Communications (2022)
-
Genomic landscape of lung adenocarcinoma in East Asians
Nature Genetics (2020)
-
Multifocal gastric adenocarcinoma in a patient with LRBA deficiency
Orphanet Journal of Rare Diseases (2017)
-
The BEACH Protein LRBA Promotes the Localization of the Heterotrimeric G-protein Golf to Olfactory Cilia
Scientific Reports (2017)